|
Using summary can give you rough snapshots of each column, but you would likely use mean, min, max, and quantile when necessary (and number of NAs): summary(tb) country 1990 1991 1992 If you would like to summarize all columns, you can use summarize_all and pass in a function (with other arguments): summarize_all(DATASET, FUNCTION, OTHER_FUNCTION_ARGUMENTS) # how to use summarize_all(avgs, mean, na.rm = TRUE) # A tibble: 1 x 10 Summarize the data: dplyr summarize functionĭplyr::summarize will allow you to summarize data. Mean_2006 media_2007 `median(\`2004\`, na.rm = TRUE)`ĬolMeans and rowMeans must work on all numeric data. If you don’t set a new name, it will be a messy output: tb %>% "2003" "2004" "2005" "2006" "2007" Summarize the data: dplyr summarize functionĭplyr::summarize will allow you to summarize data. # tb % rename(country = `TB incidence, all forms (per 100 000 population per year)`)Ĭolnames will show us the column names and show that country is renamed: colnames(tb) "country" "1990" "1991" "1992" "1993" "1994" "1995" Here we will read in a tibble of values from TB incidence: library(readxl) The matrixStats package has additional row* and col* functions.summary(x): for data frames, displays the quantile information.colSums(x): takes the sum of each column of x.rowSums(x): takes the sum of each row of x.colMeans(x): takes the means of each column of x.rowMeans(x): takes the means of each row of x.Mean(x) NA mean(x, na.rm = TRUE) 13.77778 quantile(x, na.rm = TRUE) 0% 25% 50% 75% 100%ġ 4 7 10 45 Data Summarization on matrices/data frames Note that many of these functions have additional inputs regarding missing data, typically requiring the na.rm argument (“remove NAs”). T.test will be covered more in detail later, gives a mean and 95% CI: t.test(jhu_cars$wt)ġ 3.22 18.6 2.26e-18 31 2.86 3.57 One Samp… two.sided Statistical summarization The head command displays the first 6 (default) rows of an object: library(jhur) We can use the jhu_cars to explore different ways of summarizing data. all have a na.rm for missing data - discussed later.quantile(x): displays sample quantiles of x.sd(x): takes the standard deviation of x. 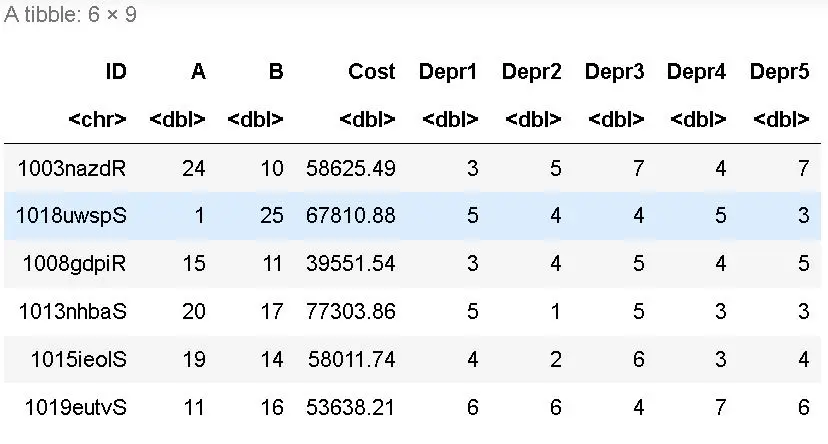
The database connections essentially remove that limitation in that you can have a database of many 100s GB, conduct queries on it directly and pull back just what you need for analysis in R. This addresses a common problem with R in that all operations are conducted in memory and thus the amount of data you can work with is limited by available memory. The benefits of doing this are that the data can be managed natively in a relational database, queries can be conducted on that database, and only the results of the query returned. An additional feature is the ability to work with data stored directly in an external database. dplyr addresses this by porting much of the computation to C . The thinking behind it was largely inspired by the package plyr which has been in use for some time but suffered from being slow in some cases. 
It is built to work directly with data frames. 
The package dplyr is a fairly new (2014) package that tries to provide easy tools for the most common data manipulation tasks.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
